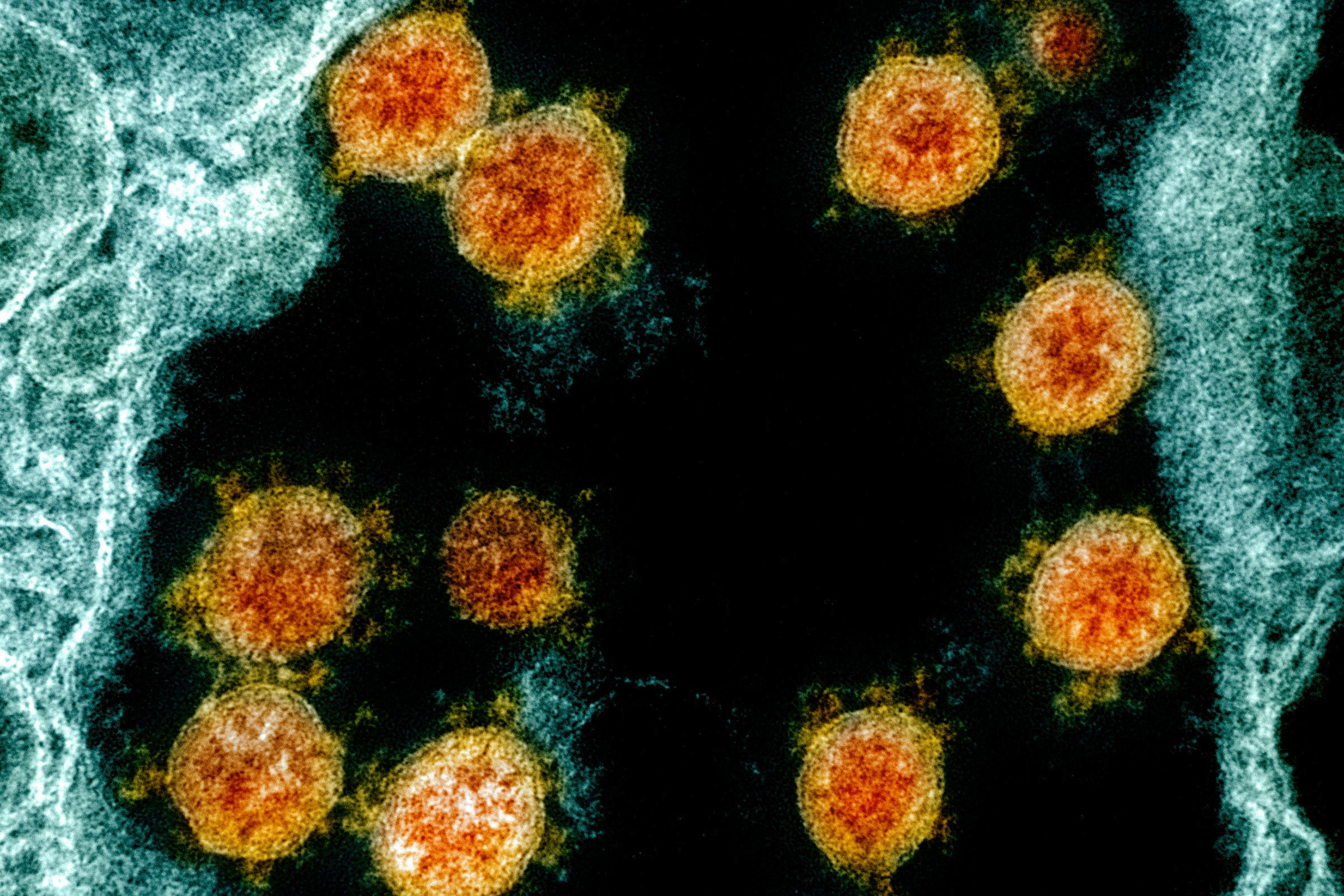
As the coronavirus pandemic continues, concern about new strains of the virus is growing. Two cases of a more contagious variant first identified in the United Kingdom have been found in Wisconsin so far.
Four labs in Wisconsin are working to sequence the genomes of virus samples from around the state to identify and track the spread of new variants.
Wisconsin’s state hygiene lab is sequencing 200 to 300 samples per week, said Allen Bateman, director of the communicable diseases division.
News with a little more humanity
WPR’s “Wisconsin Today” newsletter keeps you connected to the state you love without feeling overwhelmed. No paywall. No agenda. No corporate filter.
He says they’re looking for strains like the ones first discovered in the U.K., South Africa and Brazil.
“We all pretty much suspect that these aren’t going to be the last three variants of concern,” he said.
The Milwaukee Public Health Department, University of Wisconsin-Madison, and Gundersen Health System are also processing virus samples. Together, the four labs are sequencing an estimated 400 to 600 samples per week, Bateman said, adding he would like to see capacity increase.
In addition to identifying new strains of the virus, genomic sequencing is a way to ensure diagnostic tests are correctly identifying positive cases. PCR tests look at specific sections of the virus, Bateman said. If a mutation were to change those sections, it could lead to false negative results.
The genomic sequencing process takes longer than a PCR test, usually four or five days at the state lab. Whole genome results generally aren’t used for individual patient care or decision-making, Bateman said.
“Really what we’re doing is more in the realm of surveillance and overall public health, situational awareness,” he said.
Bateman said he hopes vaccination and other mitigation efforts will be able to slow the spread of any dangerous variants.
“It’s kind of a race now, variants versus vaccines,” he said. “How quickly can the vaccinations get out compared to how quickly can the more and more infectious variants transmit.”
Two researchers at UW-Madison began sequencing SARS-CoV-2 samples in February 2020. Virology professor Tom Friedrich and pathology professor Madison Dave O’Connor have a background in HIV research, and began sequencing SARS-CoV-2 samples from around Dane County as soon as local spread began.
“The sort of architecture of how the virus looks at the genetic level is a little different,” O’Connor said. “But the basic principles are the same as for HIV, and flu and other viruses.”
Now, their labs are processing at least 200 samples per week, and their results are published in a global database.
They hope to increase the capacity for genomic sequencing of coronavirus samples in the near future. More funding will be necessary, Friedrich said, as the supplies to process a single viral genome can cost about $50.
Wisconsin Democratic U.S. Sen. Tammy Baldwin supported measures that would dedicate $1.75 billion to genomic sequencing of virus samples, which passed a House committee last week.
Wisconsin Public Radio, © Copyright 2026, Board of Regents of the University of Wisconsin System and Wisconsin Educational Communications Board.